The ME SPUR Experience: Beattie researches image-based elastography in the deforming cell nucleus

Mechanical engineering undergraduate researcher, Julia Beattie.
The ME SPUR Program, modeled after CU Summer Program for Undergraduate Research, enabled undergraduate students to work with mechanical engineering faculty during summer 2020 on research that could be conducted remotely. As a participant, Julia Beattie worked with Professor Corey Neu to measure intranuclear mechanics. The goal was to provide a non-invasive framework to investigate the mechanobiological function of subcellular and subnuclear domains limited only by the spatiotemporal resolution of the image acquisition method. Her summer research project was titled, Image-Based Elastography of Heterochromatin and Euchromatin Domains in the Deforming Cell Nucleus.
Beattie is a third-year student from Centennial whose area of focus in the mechanical engineering department is in biomedical engineering. Beattie hopes to use her engineering education to work on medical devices or research to improve people’s quality of life. Her insights below provide a window into her research experience with ME SPUR.
Describe your summer research.
The goal of my project was to assist the Neu Lab in automating their deformation microscopy code and also, to beta test their new graphical user interface (GUI). Deformation microscopy is a technique that utilizes conventional imaging and a hyperelastic warping algorithm to analyze deformation and strain patterns within cell nuclei. In the GUI, the user inputs images of a cell nucleus before and after a mechanical deformation. Then, the program processes the images, runs a warping algorithm and creates a map of intranuclear strains. I have been running images through the program to look for errors and ways to improve it.
We made progress towards an automated, user-friendly program. The goal is for the program to be published for use in other labs where it will hopefully be used to contribute to further research of cell mechanobiology. The Neu Lab also plans to continue using deformation microscopy in future work in intracellular elastography which could potentially help in many mechanobiological applications.
What was it like conducting research remotely?
It was surprisingly uncomplicated to work in a remote format. I did most of the work on a remote desktop that I accessed from my computer. I had weekly Zoom meetings with my supervisor and other lab members. Between meetings, I regularly communicated and shared my findings with the other lab members via email and Slack.
What challenges did you encounter and work through as part of your project?
One of the main challenges was working with a Mac while the rest of the lab had PCs. Certain software packages function differently (or not at all) on my computer compared to theirs. However, the remote desktop helped, and this obstacle ultimately turned into an opportunity: my ability to run the deformation microscopy code on my computer allowed the team to start developing code that is compatible with macOS.
What about this project was rewarding?
It was very rewarding to see my findings and suggestions implemented. This program will someday be used in a variety of mechanobiological research labs, and I can say I helped make it usable.
Did you have any research experience prior to ME SPUR?
I had no experience with research prior to this program. My experience with MATLAB from past classes helped me understand the program I was working with. Additionally, my modest knowledge of cellular biology from past courses (I used to be a chemical and biological engineering major) helped me grasp the biological concepts of cell mechanics more quickly.
What advice would you share with other students considering getting involved in research?
I would highly recommend applying, even if you feel like your qualifications are not good enough. I almost didn’t apply for that reason, but I’m glad that I did. You will learn technical skills, develop communication and teamwork skills and make new connections.
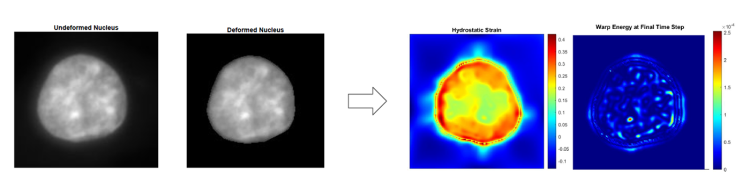
Images from Beattie's research showing an analysis of deformation and strain patterns within a cell nucleus. Inputs are shown in the left two images and results are shown in the right two images.

