Home
Welcome to Park lab!
We study the proteasome, a central mediator of protein breakdown. Functions and mechanisms of the proteasome are extremely conserved in eukaryotes, allowing our research findings to be broadly applicable to many organismal contexts, from a simple model as budding yeasts to a more complex, human stem cells, for example. A major research question in Park lab is to understand how the proteasome, a complex molecular machine, assembles via multiple dedicated chaperone proteins. How do these chaperones enable proteasome assembly accurately and efficiently? How do they 'know' faulty vs. correct complexes, and act on them properly? Cell's ability to supply the proteasome is essential to life, and survival through many stressed conditions, including "nutritonal stresses". In particular, cancer cells frequently encounter nutritional stresses, and many of them alter expressions of proteasome assembly chaperones. For more detailed description of our current projects, please see Research section of our website.
We are always interested in enthusiastic individuals at any level, including Undergraduate Students, Research Assistants, Graduate Students, or Post-Doctoral Fellows. Are you interested in research opportunities in Park lab? Please, contact Soyeon (soyeon.park-1@colorado.edu).
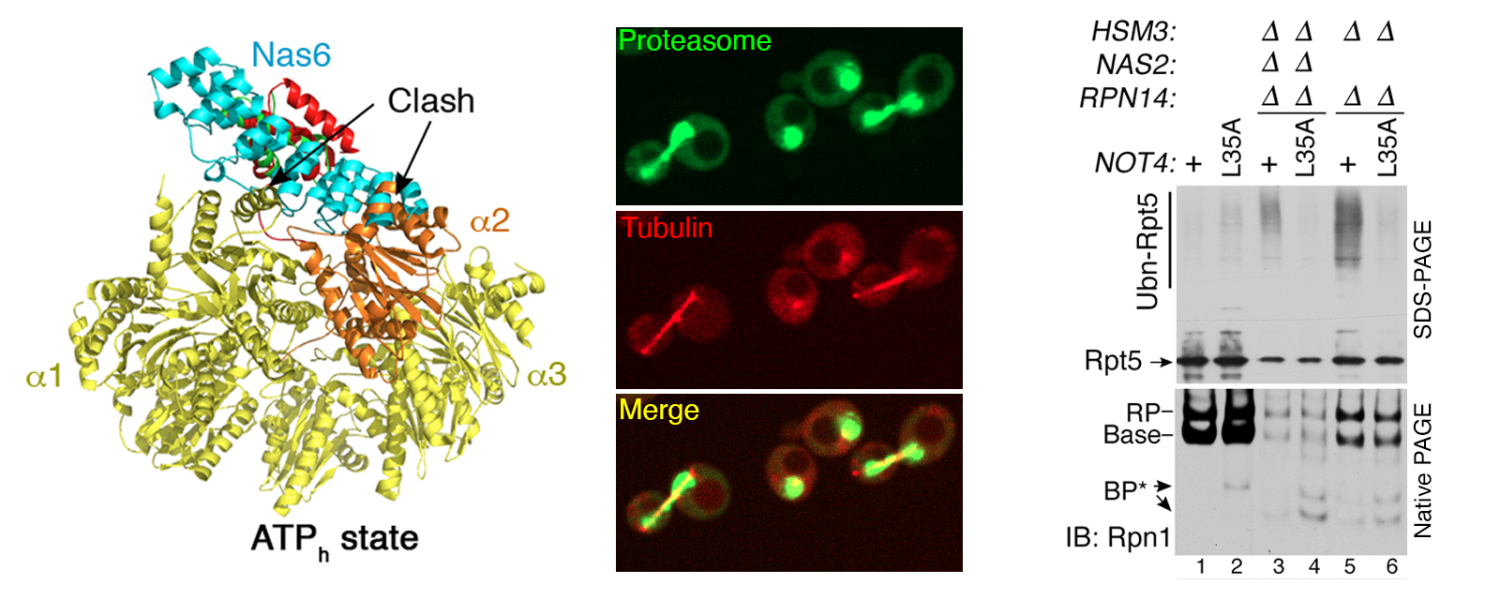
In the left panel, Nas6 (cyan) exhibits steric clash against alpha subunits (yellow) when Rpt3 (orange) is in ATP-hyrolyzing state. In the middle panel, GFP-tagged proteasome (green) and mRuby-tagged tubulin (red) are shown by live-cell confocal imaging, together with the overlay of these two images. The right panel shows that Not4 ubiquitinates Rpt5 (top) specifically when a base assembles without Nas2 and Hsm3.

