Research
Major Current Research Interests
1. Mechanisms by which micronutrients impact food uptake, foraging behaviors and development
We have established a unique system to identify essential micronutrients that are provided by live bacteria for normal C. elegans development. We are analyzing (1) how animals respond to deficiency of specific micronutrients and (2) how specific micronutrients impact cellular processes in animals. You can read more about our work that identified a vitamin B2 sensing mechanism that regulates gut protease activity to impact animal’s food behavior and growth (Qi et al. 2017 doi:10.7554/eLife.26243).
Our microbiota study has revealed an unexpected role for Enterobactin (Ent), a siderophore made by E. coli, in aiding the uptake of iron by C. elegans. We also found this role is conserved in human cells. These findings suggest that Ent from microbiota can benefit the host's iron homeostasis, and may be a useful therapy for iron deficiency (Qi and Han 2018). PDF You can read more about this work in this Cell Preview (PDF) or hear perspectives on its clinical implications from the New England Journal of Medicine (IronWars - The Host Strikes Back.) We also found that this FeEnt benefit is independent of Divalent Metal Transporter 1 (DMT1) (Sewell et al 2025, DOI: 10.1016/j.jbc.2025.108158), which may suggest a unique uptake mechanism to be explored further.
Media coverage: E. coli doesn't just give you food poisoning—it can also help you get ... CU Boulder researchers discover potential health benefit in E. coli.
Microbial Cell microreview: PDF
2. Discovery of a beneficial peptidoglycan muropeptide from E. coli that acts as an ATP synthase agonist to benefit the host.
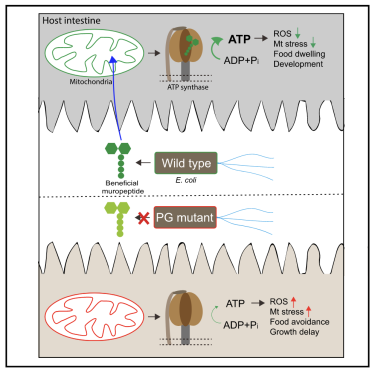
Commensal bacteria are proving to be a rich resource to the host. Our new study (Tian and Han 2022 DOI: 10.1016/j.devcel.2021.12.016) revealed a benficial role for components of the E. coli cell wall, peptidoglycan (PG) muropeptides, in promoting development and food behaviors in C. elegans. The benefit is realized when muropeptides enter the host intestine, reach the mitochondria, and bind to ATP synthase. This action prevents oxidative stress and promotes mitochondrial homeostasis. We have found that PG muropeptides function as an ATP agonist, perhaps the first ever reported.
We recently demonstrated that this unique benefit is conserved in mammals (Tian et al 2024, 10.1016/j.celrep.2024.114067), which may indicate many beneficial and therapeutic applications.
3. Mechanisms by which straight-chain fatty acids are being sensed to regulate sex-determination, reproductivity and other aspects of development.
Fat metabolism has been linked to fertility and reproductive adaptation in animals and humans, but cellular and molecular mechanisms involved in the response of reproductive development to fat levels, including a potential role of fat in environmental sex-determination, remain to be explored. Hypothesizing that a signaling mechanism perceives and interprets lipid cues to regulate the germ cell fate, we launched a study of this fatty-acid sensing mechanism that regulates germline sex-determination (Tang and Han 2017 doi:10.1016/j.cell.2017.03.049). We first observed an impact of fatty acid (FA) synthesis and mobilization by lipolysis in somatic tissues on oocyte-fate in Caenorhabditis elegans. We then identified ACS-4, an acyl-CoA synthetase and its FA-CoA product, as key germline factors that mediate the role of FA in promoting oocyte fate through protein myristoylation. Our data indicated that ACS-4-dependent protein myristoylation perceives and translates the FA level into regulatory cues that modulate the activities of MPK-1/MAPK and key factors in the germline sex-determination pathway. Our findings, including a similar role of ACS-4 in a male/female species, uncover a likely conserved mechanism by which FA, an environmental factor, regulates sex determination and reproductive development. Currently, we are carrying out projects to analyze the function of lipidation in mice and in other developmental processes in worms.
Further work identified a role for fatty acids in sarcomere integrity (Tang et al. 2021 doi: 10.1016/j.celrep.2021.109539). We found that disrupting fatty acid biosynthesis, or blocking protein myristoylation, caused muscle defects and paralysis in C. elegans. The mechanism includes ER stress and the ER unfolded protein response, with a critical role for sarcomere component PINCH as a downstream event.
4. Mechanisms by which the TORC1 signaling system senses the level of lipids to regulate postembryonic development and foraging behaviors.
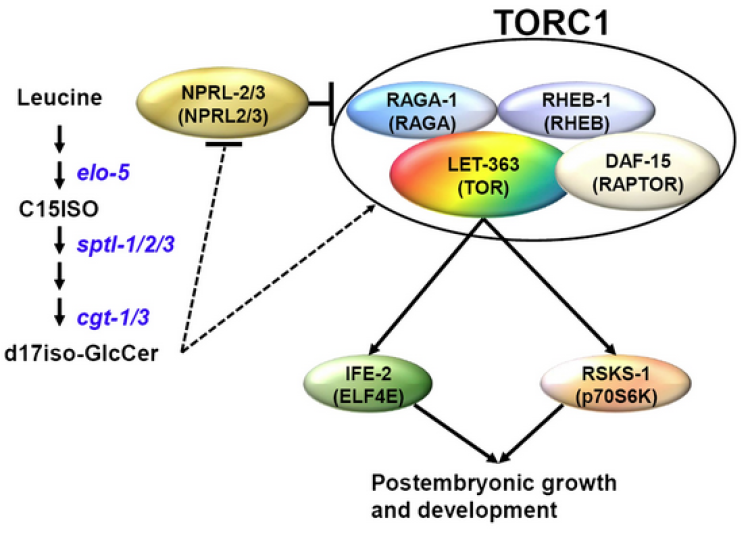
Regulation of animal development and behavior in response to nutritional cues is an intensely studied problem related to disease and aging. The existence of signaling systems that function as fatty acid (FA) or lipid sensors were unclear when we stepped into the field. We found that worms halt postembryonic development shortly after hatching in response to deficiency of monomethyl branched-chain fatty acids (mmBCFAs) that are derived from branched-chain amino acids (BCAA). We have published 13 papers since 2004 that dissect the mechanism of this response. Through genetic suppressor screens and biochemical analysis of lipid molecules, we discovered that intestinal TORC1 signaling pathway can respond to deficiency of mmBCFA through sensing the level of an mmBCFA-derived sphingolipid, d17iso-glucosylceramide (d17iso-GlcCer) (Zhu et al. 2013 doi:10.7554/eLife.00429). We then found that these lipid molecules promote foraging behavior and sensory neuron maturation through both TORC1-dependent and -independent mechanisms (Kniazeva et al. 2015 doi:10.1016/j.devcel.2015.02.015). Further study indicated that intestinal apical polarity mediates the regulation of TORC1 by GlcCer (Zhu et al. 2015 doi:10.1101/gad.263483.115). These results presented a mechanism by which lipid molecules promote development and neuronal functions involved in foraging and add critical insight into physiological functions of TORC1. While a former postdoc is continuing the study in his own lab, we have focused our study on searching for factors acting downstream of TORC1 for specific physiological functions, using both genetics (gene expression analysis and screen) and biochemistry (mass spec analysis of putative TORC1 targets). Our recent phosphoproteomic study has revealed exciting new TORC1 targets (Sewell et al. 2022 doi: 10.1016/j.isci.2022.104186).
For an extensive review of TORC1 and TORC2 studies in C. elegans, check out our WormBook chapter, TOR Signaling in Caenorhabditis elegans Development, Metabolism, and Aging.
5. Mechanisms by which animals sense the level of pyrimidines to regulate germline stem cell proliferation.
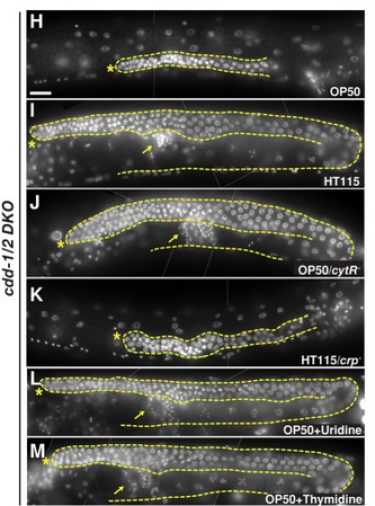
The connections between nutrient sensing systems and reproductive programs are underexplored and whether there is a mechanism that senses nucleotide (NT) levels to coordinate germline proliferation was a highly attractive problem for us a few years ago. We have established a model system where NT metabolism is perturbed both in the nematode C. elegans (cytidine deaminases, CDD) and its food (E. coli); when fed food with low uridine/thymidine (U/T) levels, germline proliferation is arrested. We provide evidence that this impact of U/T level on the germline is critically mediated by the GLP-1/Notch signaling pathway known to promote mitotic proliferation (Chi et al. 2016 doi:10.1101/gad.275107.115). Our results reveal a previously unknown NT sensing mechanism for controlling reproductivity. By a genetic suppressor screen, we isolated three mutations that significantly recovered fertility of cdd(-) mutants fed low U/T food. Cloning and analyses of these mutants will lead to mechanistic insight regarding how a low pyrimidine level is initially sensed in the intestine and the connection of this sensing system to GLP-1 activity.
You can read more about this study in this mini-review (Feeding the germline) and Editor's Choice commentary (Notch for nucleotide sensing).
Our recent follow-up study (Jia et al. 2020 doi: https://doi.org/10.1016/j.celrep.2020.01.050) revealed the role of endonuclease ENDU-2 in regulating nucleotide metabolism and germ cell proliferation. ENDU-2 responds to nucleotide imbalance and genotoxic stress, with impacts on mitotic chromosomal segregation and lifespan, to inhibit phosphorylation of CTPS-1 through repression of histone deacetylase HDA-1 and the PKA pathway. This work uncovered a new role for ENDU-2 that may lead to novel cancer therapies.

