Our research, which is highly multidisciplinary, is rooted in an interest in understanding the relationship between the structure, function, and environment of proteins and enzymes. The understanding of this relationship is crucial for rationally engineering proteins and enzymes to create new therapies, methods for the detection and destruction of toxic and/or harmful agents, and routes for the synthesis of industrially important chemicals and renewable fuels. We are particularly interested in exploiting the rich and diverse range of properties of polymeric materials to tune the activity and stability of enzymes. The current projects in the groups include:
Exploiting Enzymes in Non-Native Solvent Environments for Biocatalysis
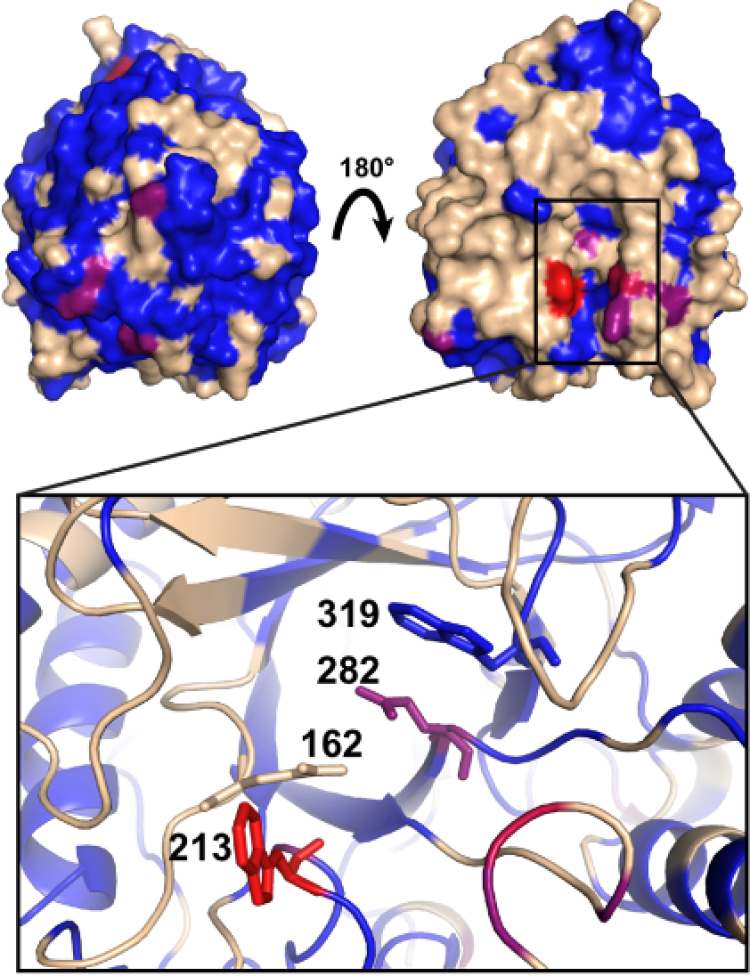
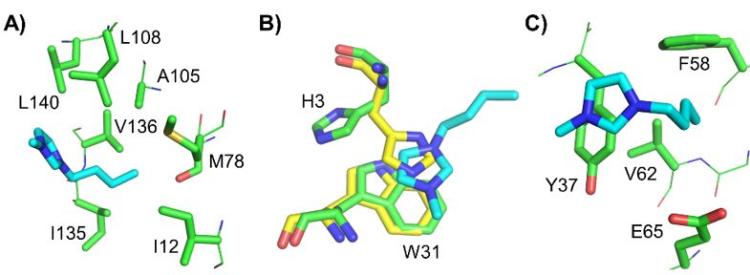
Understanding the structure and dynamics of enzymes in their solvent microenvironment is key to fully exploiting their activity. Replacing water, the natural milieu of enzymes, with anhydrous solvents can modulate, and in some instances improve, their catalytic properties. The removal of bulk water permits enzymes to catalyze very different reactions, which otherwise would not occur, alters enzyme specificity, enhances the solubility of hydrophobic reaction substrates, and suppresses undesirable hydrolytic side reactions. Additionally, enzymes are more resistant to thermal inactivation in anhydrous solvents. Consequently, the portfolio of feasible biotransformations can be greatly expanded by studying enzymes in non-natural environments. Owing to their unique and diverse molecular features, one such environment in which enzymes have been extensively studied is ionic liquids (ILs), which are comprised of solely ions. The diversity of ILs permits ILs to be rationally tailored to dissolve a broad range of reaction substrates, including cellulose and other normally recalcitrant materials. A major goal of this project is to understand how ILs impact enzyme structure and function at the molecular level as well as develop strategies to engineer enzymes for improved activity in ILs. Currently, such understanding is being used to rationally design enzymes for enhanced activity in ILs for industrial biocatalysis and bioremediation applications.
Developing and Applying Single-Molecule Methods to Elucidate the Mechanisms of Protein Interactions with Surfaces
Understanding th
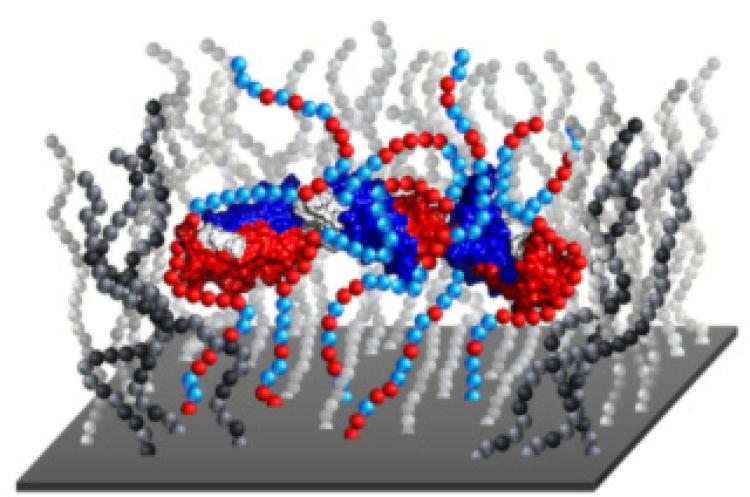
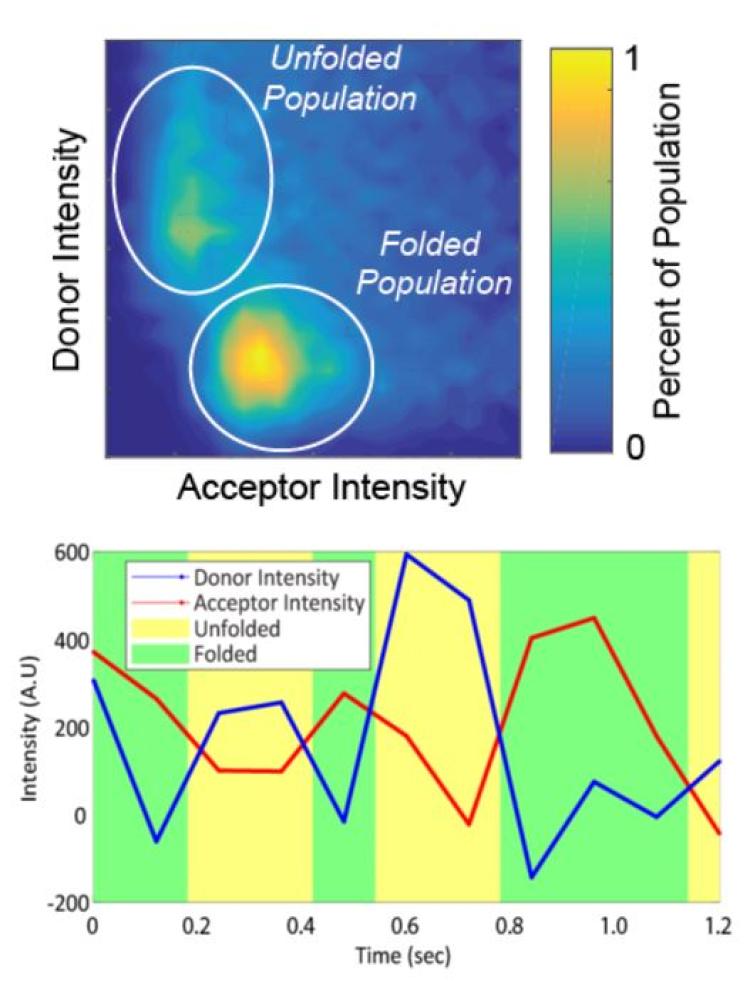
e behavior of proteins at material interfaces has become increasingly important as the intersection between the biological and synthetic worlds continues to grow. By inducing changes in protein structure, interactions with materials may lead to a loss of function and/or biological activity, as well as undesirable cellular interactions
in vivo, including the foreign body reaction to implantable materials. Despite extensive research, the understanding of the mechanisms that lead to protein conformational changes on surfaces remains incomplete. In light of this lack of understanding, we have developed novel single-molecule methods that are uniquely sensitive to monitoring protein structure and dynamics (e.g., re-folding) on surfaces. The basis for such methods involve exploiting single-molecule tracking with total internal reflection fluorescence (TIRF) microscopy in combination with Förster resonance energy transfer (FRET). Having used such methods to provide newfound insight into the mechanisms of protein unfolding on surfaces, we are now exploiting this insight to develop novel materials that stabilize proteins. Specifically, we applying this insight to the design of novel biomaterials that control protein and cellular interactions as well as immobilization supports for enzymes for biosensing applications, including the detection of chemical signatures of nuclear proliferation.
Rational Synthesis of Protein-Polymer Conjugates and Materials
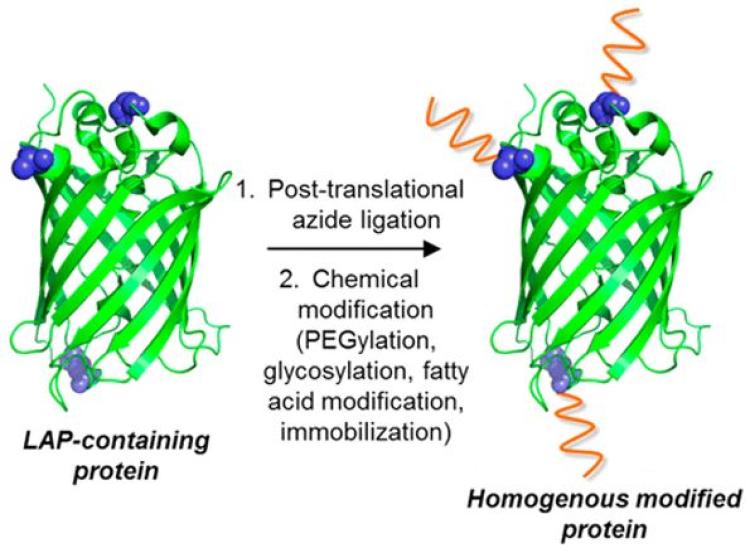
Combining proteins with polymers presents considerable opportunities to impart materials with novel and sophisticated biological function as well as tailor the activity and stability of proteins. Protein-containing polymers can be prepared by reacting a prepolymer with a protein, resulting in multipoint covalent immobilization of the protein within the polymer network. In this way, the protein is reminiscent of a monomer in the polymerization reaction. The protein is retained in the biopolymer as a result of covalent conjugation of the protein to polymer. In addition to introducing functionality, the incorporation of proteins into materials generally renders the protein more stable compared to its free state. We have recently used this approach to develop enzyme-containing coatings that inhibit for the formation of biofilms for use on medical devices (e.g., catheters). In this work, the quorum quenching enzyme acylase was specifically immobilized within biomedical grade polyurethane films. Of equal interest is covalently tethering polymers to enzymes to enhance their properties in a broad range of conditions, thus generating more robust and useful biocatalysts. While proteins have been commonly modified with PEG to improve their stability, the use of more diverse polymers has led to intriguing possibilities. In recent work, we have modified enzymes with stimuli-responsive polymers that permit the interactions of enzymes with non-aqueous solvents to be predictably and reversibly tuned. As part of this work, we are also developing bioorthogonal approaches to control the location of polymer attachment to enzymes. Through the use of such approaches, we are interested in understanding how polymer chemistry and location of attachment to enzymes impacts enzyme structure, function, and dynamics.
Combining Optically-Diffracting and Stimuli-Responsive Materials to Create a Reagent-Free Biosensing Platform
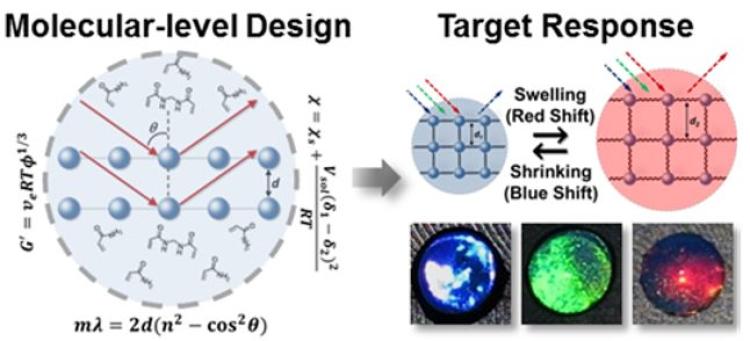
Analyte responsive hydrogels in which photonic crystals are embedded (i.e., photonic crystal hydrogels) have gained attention as a simple yet powerful platform for chemical and biological sensing. As sensors, these materials are highly sensitive to changes in the dimensions of the embedded photonic crystal over nanometer-length scales in response to an analyte. The analyte may induce swelling or contraction of the hydrogel, thereby altering the diffraction of light, and when tuned to visible wavelengths, it may elicit a change in the structural color of the hydrogel. To date, photonic crystal hydrogels have been used to detect a myriad of analytes, ranging from small molecules and organic solvents to nucleic acids and enzymes. Having demonstrated the detection of a broad range of molecules, we are currently investigating the utility of this approach as a diagnostic tool for screening for viral enzymes. Specifically, we currently have a project in which we are adapting this approach to screen for the NS2B-NS3 protease from the Zika virus. Notably, the development of affordable, sensitive, and robust methods for screening for Zika is an area of urgent need to combat the global spread of Zika. We are also interested in extending this technique for the screening of a broad range of related viruses, including HIV, West Nile, Dengue, Yellow fever, and Japanese encephalitis.