Research
Overview
Development is extraordinary feat in complex organisms. Numerous cell types with highly specific functions arise from a single cell, the zygote. During this process, the embryo must strike a fine balance between replenishing stem cell populations (i.e., self-renewal) and appropriate induction of specialized cells (differentiation). Remarkably, all of these decisions take place largely without changing the genomic content of the cells involved, which underscores the importance of intricate and dynamic regulatory mechanisms in development. Understanding these mechanisms is crucial for both basic science and medicine. Our lab seeks to develop and apply novel and innovative tools to study the molecular determinants of cell fate.
Reprogramming
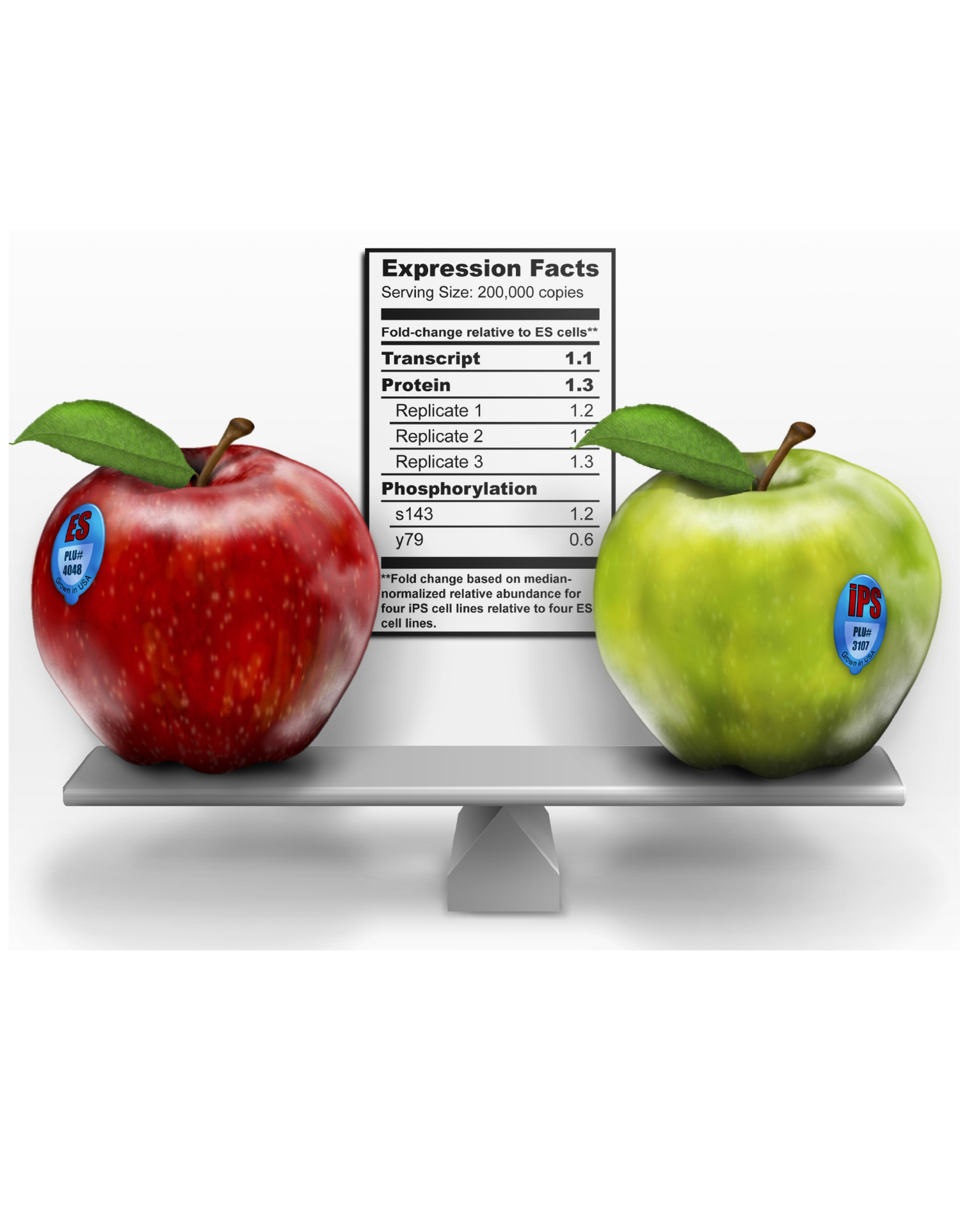

For a long time, it was thought that cell fate change was unidirectional. In other words, stem cells progressively differentiate, lose plasticity, and specialized progeny are restricted to their respective cell type. We know now that fully differentiated cells can be experimentally induced to pluripotency through a process called reprogramming. Comprehensive analyses have shown that these so-called induced pluiripotent stem cells (iPS cells) are molecularly and functionally highly similar to embryonic stem (ES) cells. Moreover, somatic cells can transition to entirely different somatic cell types (e.g., fibroblasts into neurons) through transdifferentiation. These experimental approaches provide exciting opportunities in regenerative medicine and drug testing; however, they are also tractable systems to study cell fate change. Our lab harnesses this power to understand the epigenetic and post-transcriptional mechanisms that instruct cell identity.
Chromatin
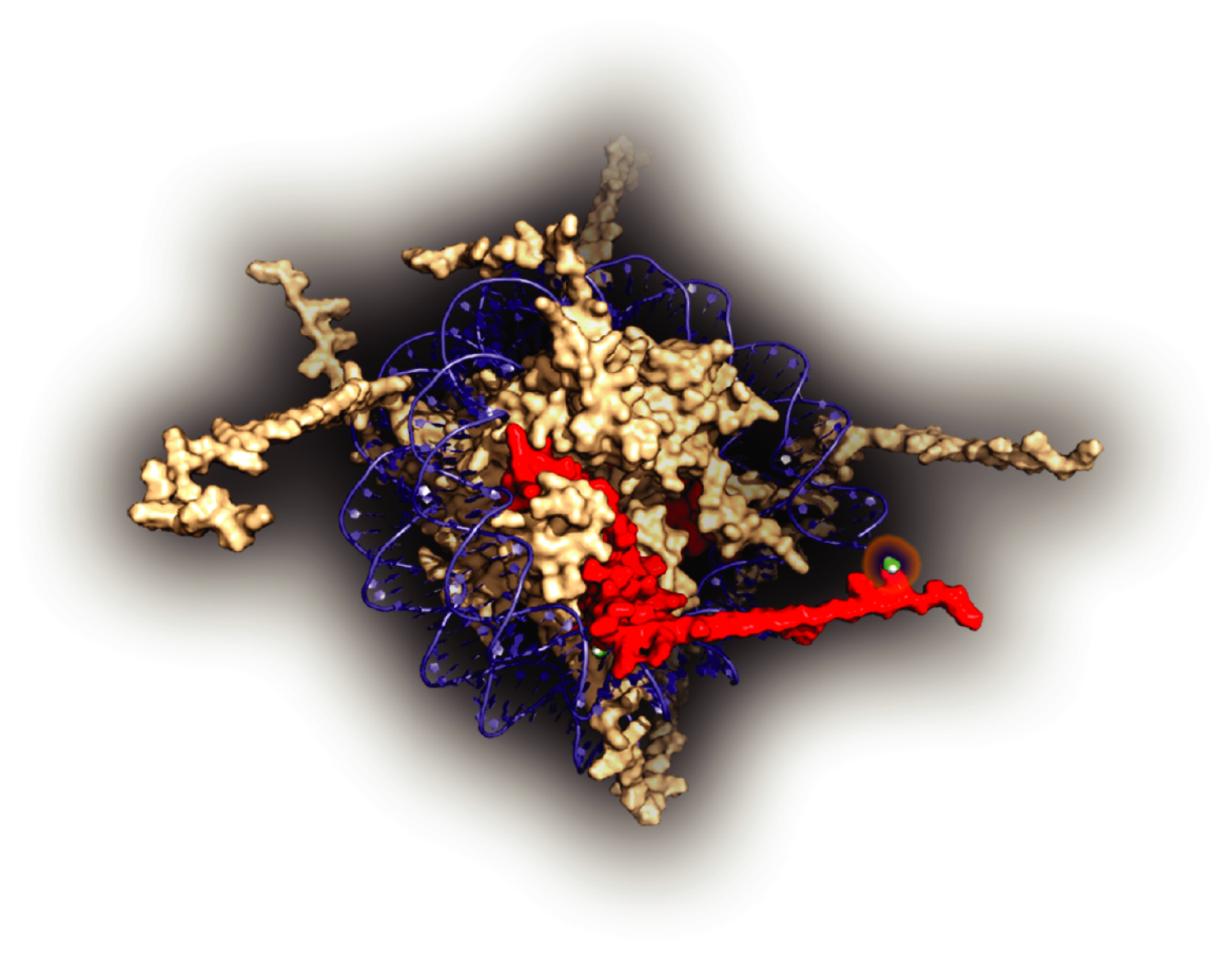
Histones are core components of chromatin and key regulators of many aspects of cell biology. The expansive roles reflect the large number of post-translational modifications that decorate histone tails. Although modification of specific residues has been studied extensively in the context of transcription, it is challenging to manipulate histone marks during biologically relevant cell fate transitions. We have developed systems to explore the role of histone methylation at specific sites on histone H3. Our lab applies this system to determine the role of chromatin modification during development and homeostasis. (pdb reference for the image: 1AOI)
RNA processing
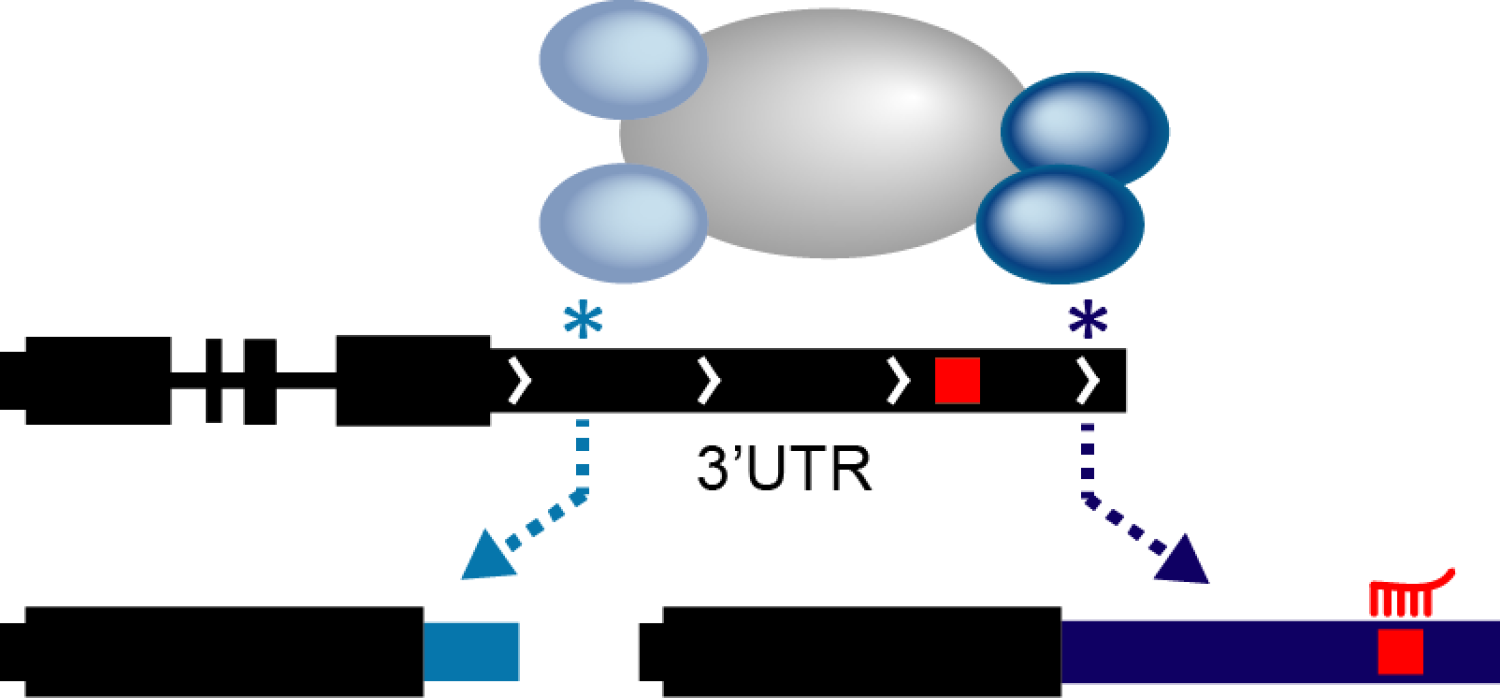
Development requires rapid and fine-tuned regulation of gene expression patterns. RNA processing is a regulatory mechanism that meets both of these needs and recently alternative polyadenylation (APA) has emerged as a widespread form of RNA processing that controls cell fate change. APA effectively controls the length of the 3’ UTR for a given transcript, thereby including or excluding regulatory sequences (e.g., miRNA seed sequences). Although it is currently known that global 3’ UTR length changes during development, it is not clear how this phenomenon relates to differentiation and cell fate transitions. Our lab is exploring the proteins that control APA and investigating their role in stem cell function.

