Unraveling the biological roles of specific miRNAs, from experimental target identification through functional characterization
microRNAs (miRNAs) are essential regulators of gene expression in diverse biological processes and disease states. They are short 22-nt noncoding RNAs that bind to mRNA targets via partial base complementarity to downregulate gene expression. Critical for understanding how any miRNA functions is identifying the set of mRNA targets to which it binds in a specific biological context, and also determining how these interactions regulate cellular processes to ultimately impact phenotype. A single miRNA has the potential to target thousands of different mRNAs. Hence, elucidating which mRNAs are bona fide functional targets of a miRNA in a specific cellular context is challenging.
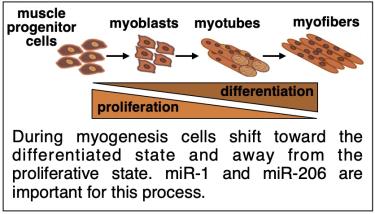
We are using differentiation of skeletal muscle cells (myogenesis) as a model system to understand miRNA-mediated regulation of gene expression. The myogenic differentiation program is driven by changes in gene expression. Part of this program involves post-transcriptional control by miRNAs known as myomiRs. Our research primarily focuses on two myomiRs, miR-1 and miR-206, that are critical for controlling myogenesis. Interestingly, these miRNAs are highly similar in sequence, yet they have distinct roles.
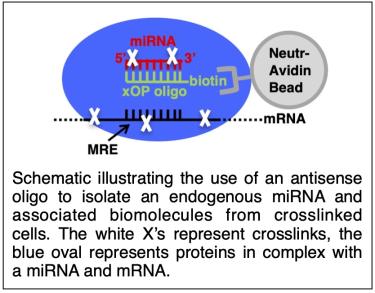
We are developing a comprehensive experimental strategy to unravel the distinct roles of miR-1 and miR-206 during myogenesis. This includes experimental target identification, screens to identify phenotypically important targets, and experiments to reveal mechanisms of regulation. The ultimate goal is to develop a general strategy to investigate the function of any given miRNA in diverse biological systems. Central to our work is an affinity purification approach to purify specific miRNA/mRNA/protein complexes and identify the mRNAs via high-throughput sequencing.

