Modeling spatiotemporally varying protein–protein interactions in CyLaKS, the Cytoskeleton Lattice-based Kinetic Simulator
Shane A. Fiorenza, Daniel G. Steckhahn,and Meredith D. Betterton (2021). European Physical Journal E 44,105. DOI: 10.1140/epje/s10189-021-00097-8. BioRxiv link. Download.
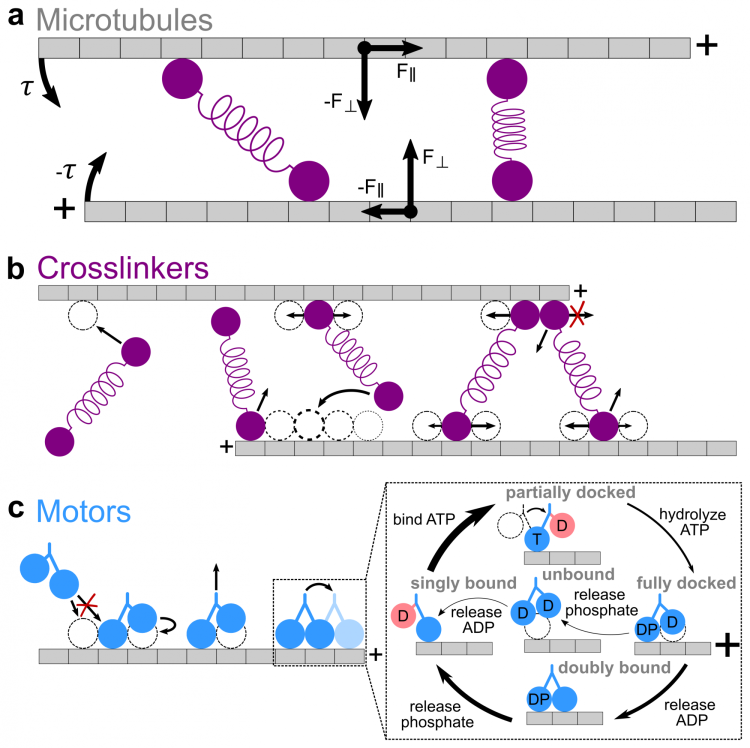
Interaction of cytoskeletal filaments, motor proteins, and crosslinking proteins drives important cellular processes such as cell division and cell movement. Cytoskeletal networks also exhibit nonequilibrium self-assembly in reconstituted systems. An emerging problem in cytoskeletal modeling and simulation is spatiotemporal alteration of the dynamics of filaments, motors, and associated proteins. This can occur due to motor crowding, obstacles along the filament, motor interactions and direction switching, and changes, defects, or heterogeneity in the filament binding lattice. How such spatiotemporally varying cytoskeletal filaments and motor interactions affect their collective properties is not fully understood. We developed the Cytoskeleton Lattice-based Kinetic Simulator (CyLaKS) to investigate such problems. The simulation model builds on previous work by incorporating motor mechanochemistry into a simulation with many interacting motors and/or associated proteins on a discretized lattice. CyLaKS also includes detailed balance in binding kinetics, movement, and lattice heterogeneity. The simulation framework is flexible and extensible for future modeling work and is available on GitHub for others to freely use or build upon. Here we illustrate the use of CyLaKS to study long-range motor interactions, microtubule lattice heterogeneity, motion of a heterodimeric motor, and how changing crosslinker number affects filament separation.

