Collective motion of driven semiflexible filaments tuned by soft repulsion and stiffness
Jeffrey M. Moore, Tyler N. Thompson, Matthew A. Glaser, and Meredith D. Betterton (2020). Soft Matter16, 9436. DOI: 10.1039/D0SM01036G. arXiv DOI:1909.11805. Download.
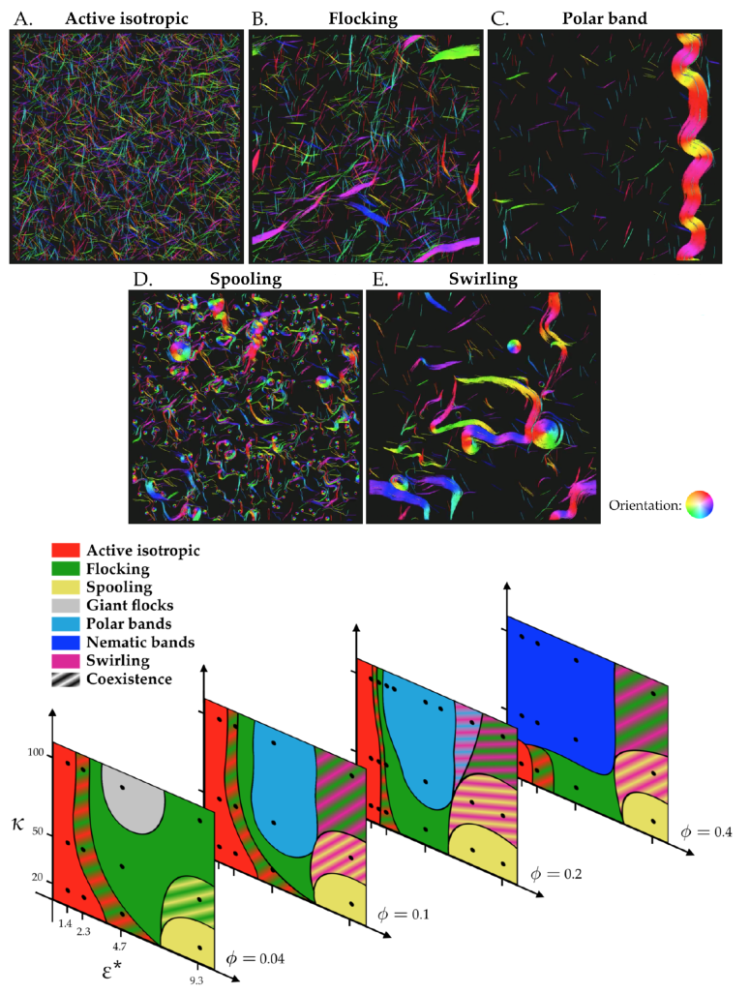
In active matter systems, self-propelled particles can self-organize to undergo collective motion, leading to persistent dynamical behavior out of equilibrium. In cells, cytoskeletal filaments and motor proteins exhibit activity and self-organization into complex structures important for cell mechanics, motility, and division. Collective dynamics of cytoskeletal systems can be reconstituted using filament gliding experiments, in which cytoskeletal filaments are propelled by surface-bound motor proteins. These experiments have observed diverse behavior, including flocks, polar streams, swirling vortices, and single filament spirals. Recent experiments with microtubules and kinesin motor proteins found that the effective repulsive interaction between filaments can be tuned by crowding agents in solution, altering the collective behavior. Adding a crowder reduced filament crossing, promoted alignment, and led to a transition from active, isotropically oriented filaments to locally aligned polar streams. These results suggest that tunable soft repulsion can control active phase behavior, but how altering steric interactions and filament stiffness alter collective motion is not fully understood. Here we use simulations of driven filaments with tunable soft repulsion and rigidity in order to better understand how the interplay between filament flexibility and steric effects can lead to different active steady states. We identify swirling flocks, polar streams, buckling bands, and spirals, and describe the physics that govern transitions between these states. In addition to repulsion, tuning filament stiffness can promote collective behavior, and controls the transition between active isotropic filaments, locally aligned flocks, and polar streams.

