Home
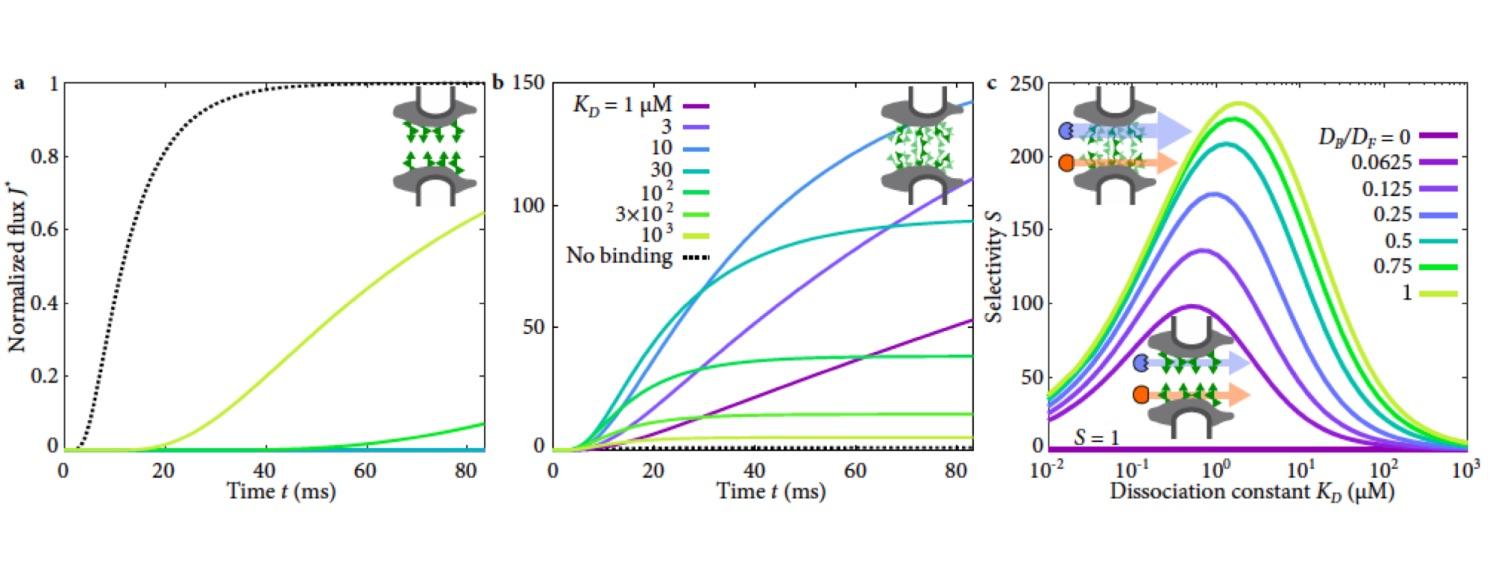
In biological systems, polymeric materials block the movement of some macromolecules while allowing the selective passage of others. We developed a model motivated by features of the nuclear pore complex (NPC) which are highly conserved and could potentially be applied to other biological systems. We show that a single feature of the NPC is sufficient for selective transport: the bound-state motion resulting from transient binding to flexible filaments. We generalized this observation to model nanoparticle transport through mucus as well. Our model provides a framework to control binding- induced selective transport in biopolymeric materials.
Welcome to the page of Betterton lab, focusing on research of Meredith Betterton and our close faculty collaborators Matthew Glaser and Richard McIntosh. We study a range of problems in molecular and cellular biophysics using experimental, theoretical, and computational approaches. Learn more about our research, papers, group members, and funding.
Our group is part of a broad biophysics and quantitative biology community at CU Boulder. If you'd like to discuss working with us, please contact Meredith Betterton.

